Author: xycarto
-
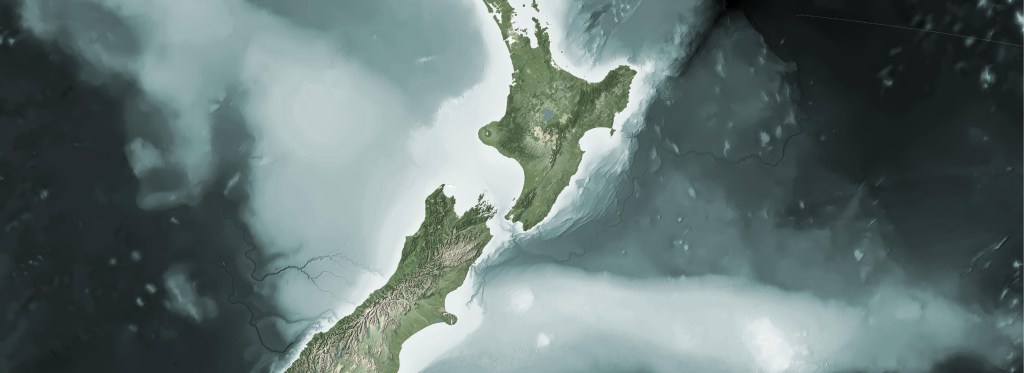
Cartographic Aotearoa
I try hard to keep this blog about basic open source processing in GIS. I don’t spend much time talking about my cartographic work, since I like put it up in the gallery and try to let it speak for itself. Further, 90% of my cartographic products involve processing data before it ever gets to…
-
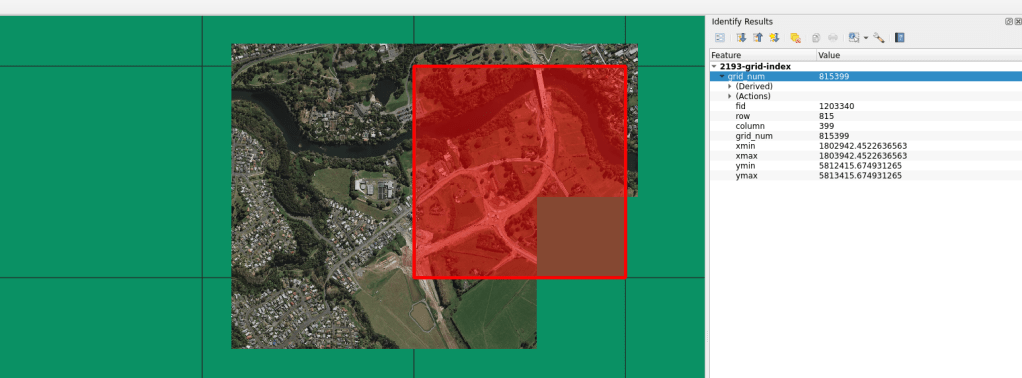
Projection Grid
There are times when I need a regular grid for an entire projection extent. Meaning, for the extent of an entire projection, I need to create a regular grid of uniform tiles across the projection. In past projects, these grids have been very helpful for data alignment and clipping data into uniform shapes and sizes.…
-

GRASS GIS, Docker, Makefile
Small example of using Docker and Makefile to implement GRASS GIS. This blog is written to be complimentary to the Github repository found here. Included in this post is a more verbose explanation of what is happening in the Github repository. Users can explore the scripts to see the underlying bits that make it each…
-

Basic COG in Openlayers: Single Band Tif
This post covers loading a raw COG Tif and manipulating the values in JS. If you are unfamiliar with the COG Tif format, see here for an explanation. Openlayers has a few good examples on how to load COGs. Some of this is a repeat of their examples and some goes a little more in…
-
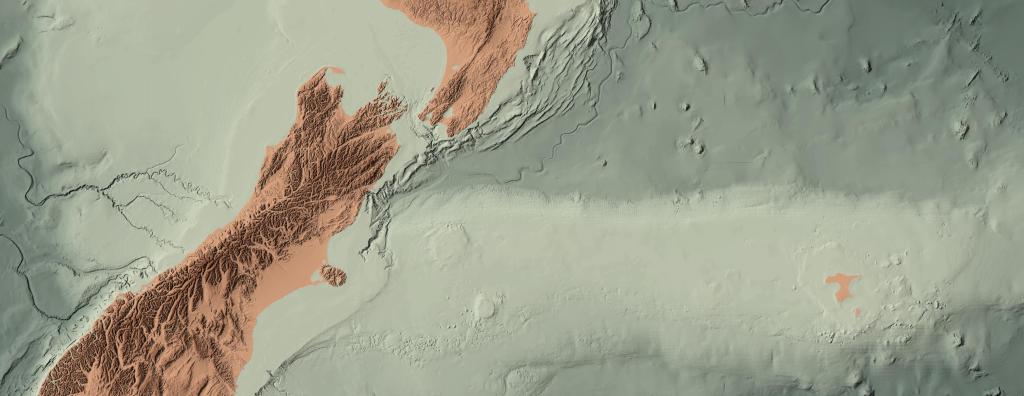
Warping Across Anti-meridian: NZTM to Web Mercator
TL;DR Python example here. Git repo updated 13 Feb 2024. NZTM to Web Mercator There are a number of times when I am building a web map when I need to warp a GTiff across the anti-meridian. New Zealand sits at the edge of this line, with a portion of it on the other side,…
-
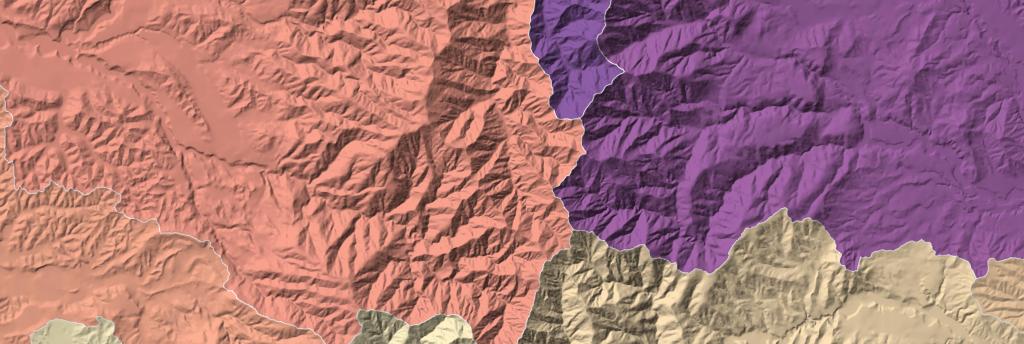
Static Vector Tiles II: Openlayers with Custom Projection
Building off the work from my previous post on vector tiles, I wanted to develop a second process for use in Openlayers. There are many tutorials out there demonstrating the use of vector tiles with predefined styles and tutorials explaining how to build a fully vector tile map online. My aim instead is to demonstrate…
-

RGB Elevation Creation for 3D Online Mapping (Terrain RGB)
The following is about how to build an elevation file to work with procedural-gl.js. I just really wanted to build my own elevation dataset and thought it would be helpful to share how I did it. Procedural-gl.js has changed the game in attainable 3D online mapping. When I came across this project, I was very…
-

Basic GRASS GIS with BASH, plus GDAL
As a follow-up to the last blog, I thought it would be helpful to demonstrate how next to break up the elevation example into individual watersheds. The reason being, in my last example I demonstrated the process using a square raster tile. Hydrological processes are not accurate when run on square tiles. It is best…
-

Basic GRASS GIS with BASH
I love GRASS… GIS But this wasn’t always the case. GRASS GIS was, for a long time, something I dismissed as ‘too complex’ for my everyday geospatial operations. I formulated any number of excuses to work around the software and could not be convinced it had practical use in my daily work. It was ‘too…
-

Wellington Elevations: Interpolating the Bathymetry
It is important to note something from the very beginning. The interpolated bathymetry developed in this project does not reflect the actual bathymetry of the Wellington Harbour. It is my best guess based on the tools I had and the data I worked with. Furthermore, this interpolation is NOT the official product of any institution.…
-

Building the Wellington Model with 1m DEM and DSM
As interest in LiDAR derived elevation increases, so grows the interest in the capabilities. LiDAR derived elevation data has been great for my visualization game and in helping me communicate the story out about what LiDAR can do. It all starts with a picture to get the imagination going. The Wellington model derived for this…
-

The Rejects
Sometimes there is simply not enough room for all the ideas. Sometimes you want all the images to make it to the final round. In a recent project to promote some of our elevation data, I was asked to present a number of ideas for a 2000mm x 900mm wall hanging. The piece was to…
-

Processing and Visualizing Auckland 1m DEM/DSM Elevation Data
About two years ago, I took on a cartographic project visualizing the Auckland 1m DEM and DSM found publicly via the LINZ Data Service (LDS) here: DEM, DSM. The goal at the time was to develop a base map for the extraction of high resolution images for use in various static media. It was a…

